Plant development and diversity
We seek to address two fundamental questions in biology: how do biological forms develop and what is the basis for their diversity. To address the first question we aim to elucidate how genotypes are translated into organismal morphologies through the process of morphogenesis. To address the second, we seek to conceptualize how the balance of conservation versus divergence in gene regulatory networks yields different morphologies during evolution. To move forward on these problems we make heavy use of genetics while also using biological imaging, genomics and computational modelling. We hope that working at the interface of these areas will allow us to understand how biological forms are produced, maintained and how they diversify.
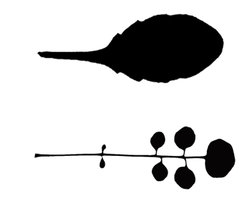
The simple serrated leaf of A. thaliana (top) vs the compound leaf of C. hirsuta (bottom).
To help our work on these problems and to facilitate an in-depth and unbiased understanding of how the balance of conservation versus divergence of development programmes results in morphological diversity, we developed the Arabidopsis thaliana relative Cardamine hirsuta into a model system for studies of evolution of development. C. hirsuta is amenable to both forward and reverse genetics approaches including transgenesis, but shows morphological divergence from A. thaliana in several key traits including leaf shape, shoot branching, floral morphogenesis and fruit development. Consequently comparative studies between these two species can greatly enrich our knowledge of the molecular mechanisms driving evolution of form. To support our comparative work we are developing extensive experimental tools for studying C. hirsuta development and diversity. These include genome assemblies of a panel of natural strains, recombinant inbred lines, mutant populations, inducible and tissue-specific gene expression systems and platforms for time-lapse analysis of gene expression and tissue growth. We believe that in-depth comparison of genetic networks of A. thaliana vs C. hirsuta, coupled with studies of natural variation within both species and broader phylogenetically informed studies across land plants, will help us understand the genetic basis for evolutionary change.
We are pursuing three interrelated strands of research
1. Morphogenesis and control of form
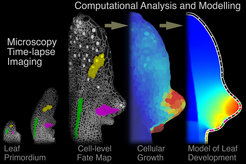
Can we conceptualize morphogenesis in a predictive fashion? The form of an organism is determined by a cascade of developmental processes that take place at different levels of organization and yield the final form through complex feedback loops of genetic regulation, signaling, and tissue growth. We aim to delineate such interactions and develop predictive models that conceptualize the process of development. Examples of the processes we are studying are leaf morphogenesis and patterning, and cell fate delimitation during embryo, shoot and root development.
2. The mechanistic basis for morphological diversity

Is morphological diversity between species generated by a large number of genetic differences of small effect, or by a few genetic changes with large effects? Are a handful of key genes responsible for evolution of multiple morphological traits? How are genes underlying diversity positioned within Genetic Regulatory Networks influencing form? Which specific genes have changed to produce the vast degree of morphological diversity seen in nature? And how do those genes change e.g. do mutations causing diversity tend to be coding or regulatory, do they behave as dominant or recessive, are they pre-existing in populations or does their sudden occurrence generate diversity?
3. Paths of evolutionary change

Natural variation in C. hirsuta leaf form. We are using genetic approaches to study the causes and consequences of such differences in form.
How repeatable is evolution? Does diversity in the same types of genes or pathways underlie variation in the same trait in different instances? Does inter- and intra-specific variation in morphology of the same traits arise via equivalent morphogenetic avenues? How prevalent is the role of positive selection in sculpting diverse plant forms and can the agents of selection be identified?
Selected Publications
- Li X-M, Jenke H, Strauss S, Wang Y, Bhatia N, Kierzkowski D, Lymbouridou R, Huijser P, Smith RS, Runions A, Tsiantis M: Age-associated growth control modifies leaf symmetry and enabled leaf shape diversification. Current Biology,34: 4547-4558 (2024).
- Hu, Z.-L.*, Wilson-Sánchez, D.*, Bhatia, N., Rast-Somssich M. I., Wu, A., Vlad, D., McGuire, L., Nikolov, L. A., Laufs, P., Gan, X., Laurent, S., Runions, A., Tsiantis, M. (2024) A CUC1/auxin genetic module links cell polarity to patterned tissue growth and leaf shape diversity in crucifer plants. Proc Natl Acad Sci (26):e2321877121. doi: 10.1073/pnas.2321877121.
- Li, X.-M., Jenke, H., Strauss, S., Bazakos, C., Mosca, G., Lymbouridou, R., Kierzkowski, D., Neumann, U., Naik, P., Huijser, P., Laurent, S., Smith, R. S., Runions, A., Tsiantis, M. (2024) Cell-cycle-linked growth reprogramming encodes developmental time into leaf morphogenesis. Current Biology, 34:3, 541-556, doi:10.1016/j.cub.2023.12.050.
- Baumgarten L*, Pieper B*, Song B, Mane S, Lempe J, Lamb J, Cooke EL, Srivastava R, Strutt S, Zanko D, Casimiro PG, Hallab A, Cartolano M, Tattersall AD, Huettel B, Filatov DA, Pavlidis PA, Neuffer B, Bazakos C, Schaefer H, Mott R, Gan X, Alonso-Blanco C, Laurent S, Tsiantis M:Pan-European study of genotypes and phenotypes in the Arabidopsis relative Cardamine hirsuta reveals how adaptation, demography, and development shape diversity patterns. PLoS Biology, 21: e3002191, (2023).
- Bhatia, N., Wilson-Sanchez, D., Strauss, S., Vuolo, F., Pieper, B., Hu, Z., Rambaud-Lavigne, L., & Tsiantis, M.(2023) Interspersed expression of CUP-SHAPED COTYLEDON2 and REDUCED COMPLEXITY shapes Cardamine hirsutacomplex leaf form. Current Biology. doi:10.1016/j.cub.2023.06.037.
- Wang Y, Strauss S, Liu S, Pieper B, Lymbouridou R, Runions A & Tsiantis M. The cellular basis for synergy between RCO and KNOX1 homeobox genes in leaf shape diversity. Current Biology, 32: 3773-3784, doi: https://doi.org/10.1016/j.cub.2022.08.020, (2022).
- Zhang, Z., Runions, A., Mentink, R. A., Kierzkowski, D., Karady, M., Hashemi, B., Huijser, P., Strauß, S., Gan, X., Ljung, K., & Tsiantis, M. (2020). A WOX/Auxin Biosynthesis Module Controls Growth to Shape Leaf Form. Current Biology, 30,4857-4868, doi:10.1016/j.cub.2020.09.037.
- Kierzkowski, D*., Runions, A.*, Vuolo, F., Strauss, S., Lymbouridou, R., Routier-Kierzkowska, A.-L., Wilson-Sánchez, D., Jenke, H., Galinha, C., Mosca, G., Zhang, Z., Canales, C., Dello Ioio, R., Huijser, P., Smith, R. S., Tsiantis, M., (2019) A Growth-Based Framework for Leaf Shape Development and Diversity. Cell, 177: 1405-1418., doi: 10.1016/j.cell.2019.05.011
* contributed equally.



