Ready, Steady, Go
Cryo-electron microscopy reveals the molecular steps in plant immune receptor activation
Two landmark studies from the laboratory of Alexander von Humboldt Professor Jijie Chai provide unprecedented structural insight into how plant immune receptors are primed – and then activated – to provide plants with resistance against microbial pathogens
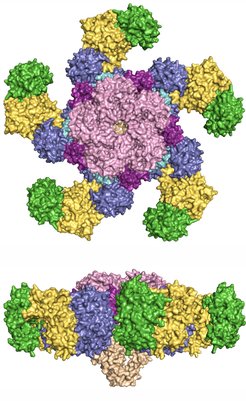
The ZAR1 resistosome.
The upper image depicts the pentameric wheel-like structure of the ZAR1 resistosome from above. In the lower panel depicting the resistosome in the side view, the funnel can be seen in light brown. PBL2 is colored in green and RKS1 in mustard. All other colors denote different domains of the ZAR1 protein.
Although separated by more than one billion years of evolution, plants and animals have hit upon similar immune strategies to protect themselves against pathogens. One important mechanism is defined by cytoplasmic receptors called NLRs that, in plants, recognize so-called effectors, molecules that invading microorganisms secrete into the plant’s cells. These recognition events can either involve direct recognition of effectors by NLRs or indirect recognition, in which the NLRs act as ‘guards’ that monitor additional host proteins or ‘guardees’ that are modified by effectors. Host recognition of effectors, whether direct or indirect, typically results in cell death to confine microbes to the site of attempted infection. However, until now, a detailed understanding of the mechanisms of action of plant NLRs has been lacking, and much of our understanding of how these molecules function in plants has been based on comparison with animal counterparts.
In two new studies published in the journal Science, Jijie Chai and his laboratory together with the groups of Hong-Wei Zhang and Jian-Min Zhou at Tsinghua University and the Chinese Academy of Sciences in Beijing have now pieced together the sequence of molecular events that convert inactive NLR molecules into active complexes that provide disease resistance.
The authors focused their attentions on a protein called ZAR1, an ancient plant molecule that is likely to be of broad importance since it is capable of indirectly sensing several unrelated bacterial effectors. For indirect pathogen recognition inside plant cells ZAR1 receptor must form a preformed complex with another plant protein, called RKS1.
Using cryo-electron microscopy, Chai and co-authors observed that in the absence of bacterial effectors, ZAR1, together with RKS1, is maintained in a latent state through interactions involving multiple domains of the ZAR1 protein. Upon infection, a bacterial effector modifies the plant ‘guardee’ PBL2, which then activates RKS1 resulting in huge conformational changes that first allow plants to swap ADP for ATP and then result in the assembly of a pentameric, wheel-like structure that the authors term the ‘ZAR1 resistosome’.
One striking feature of this structure is its similarity with animal NLR proteins, which, once activated, also assemble into wheel-like structures that act as signaling platforms for cell death execution and immune signaling. However, one important difference between the structures offers a tantalizing clue as to how ZAR1 induces cell death. The authors could identify a highly ordered funnel-like structure in ZAR1 that tethers the resistosome to the plasma membrane and is required for cell death and disease resistance. The authors speculate that ZAR1 may form a pore in the plasma membrane and in this way perturb cellular function leading to immune signaling and cell death.
Other plant NLRs also assemble into complexes that associate with the plasma membrane and it is thus highly likely that Chai’s findings have important general implications for understanding plant immunity. MPIPZ director Paul Schulze-Lefert, who was not involved in the studies, is in no doubt about the importance of the new studies: “This will become textbook knowledge.”












