Plant homeodomain proteins promote synthesis of the hormone auxin to help leaves grow wide
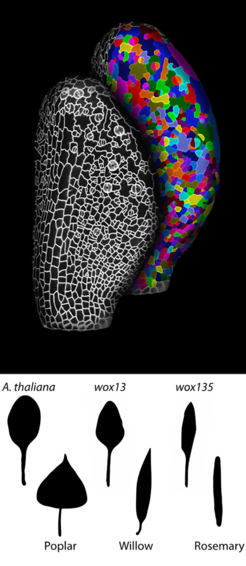
A comprehensive analysis of the development and diversity of leaf shape promises to answer one of the longest-standing questions in biology – how gene activity influences cellular growth and form to create the organ shapes of plants and animals. Recent findings presented by Dr. Zhongjuan Zhang, Dr. Miltos Tsiantis and their colleagues at the Max Planck Institute for Plant Breeding Research (Cologne, Germany) in an article published in Current Biology offer important advances in our understanding of morphological diversity using plant leaves as an example. The researchers found that WOX homeodomain transcription factors, which are key regulators of plant development, regulate leaf form by controlling the amount and direction of growth as well as the rate of differentiation. Specifically, they found that spatially restricted WOX-mediated control of biosynthesis of the hormone auxin is instrumental for lateral leaf growth and acquisition of the simple, ellipsoid leaf form observed in A. thaliana. Time-lapse imaging was utilized to measure growth at cellular resolution during development and, consequently, learn how WOX factors influence cell growth to shape leaf geometry. Analysis of this rich dataset required the generation of various fate maps and comparing the balance of conservation versus divergence in cell-level growth properties at equivalent positions in the leaf primordium in different genotype.
This approach is essentially the developmental biology equivalent of a sequence alignment. Instead of nucleotides, cell growth parameters are compared in equivalent positions in primordia of different genotypes. Notably, this methodology has the potential to become a fundamental tool in plant biology. Dr. Adam Runions, a computer scientist involved in this study, who has gone ahead to start his own research group at the University of Calgary, said “it was exciting to see how the computational methods we developed to analyse time-lapse growth data could help us understand how genetic factors control growth to shape organ form". Computational analysis of leaf imaging data was complemented with genetic testing to clarify the necessity of auxin and WOX genes for tissue level growth and patterning. Overall this analysis revealed that theWOX-auxin interaction is important for leaf width, as wox mutant plants develop narrow leaves and this effect can be partially reversed by genetically increasing auxin production in the WOX domain. Specifically, auxin biosynthetic gene expression can partially by-pass the requirement for WOX genes during development when supplied in the WOX expression domain, but not the epidermal domain. Together, these findings demonstrate for the first time that the precise spatial deployment of the WOX-auxin module is critical for leaf shape. It also provides a rare example where a downstream process, in this case, the auxin biosynthetic gene expression, is sufficient to account for a measurable component of activity of the upstream transcription factors as seen with WOX. The authors hope their work encourages additional investigations exploring how regulated growth controls organ geometry using both a theoretical and experimental approach.
Dispatch article highlighting the work:
