Plant-microbe homeostasis: a delicate balancing act
Scientists from the Max Planck Institute for Plant Breeding Research in Cologne, and the University of North Carolina at Chapel Hill, have shown that the presence of both immune-suppressive and non-suppressive bacteria in the plant root microbiota is crucial to strike a balance between plant growth and plant defence, and maintain plant-microbe homeostasis.
Plants grown in soil are colonized by diverse microbes collectively known as the plant microbiota, which is essential for optimal plant growth in nature and protects the plant host from the harmful effects of pathogenic microorganisms and insects. However, in the face of an advanced plant immune system that has evolved to recognize microbial associated-molecular patterns (MAMPs) – conserved molecules within a microbial class – and mount an immune response, it is unknown how soil-dwelling microbes are able to colonize plant roots. Now, MPIPZ researchers led by Paul Schulze-Lefert, and researchers from the University of Carolina led by Jeffery L. Dangl show, in two separate studies, that a subset of commensal bacteria is able to suppress a sector of the plant immune system and colonize plant roots when both immune-suppressive and non-suppressive bacteria are present, in the context of a microbial community.
Plants have evolved an innate immune system to protect themselves against pathogens, including the recognition of microbe-derived MAMPs by pattern recognition receptors (PRRs) that reside on the surface of plant cells. Recognition of MAMPs by PRRs leads to MAMP-triggered immunity (MTI), which results in the restriction of pathogen proliferation. However, left unchecked, chronic MTI can also lead to plant growth inhibition, a phenomenon known as growth-defence trade-off. Pathogens have evolved diverse mechanisms to supress MTI, a property that has, surprisingly, also been detected in a subset of non-harmful commensal bacteria of the root microbiota. These bacteria have been coined ‘immune-suppressive’ microbes, in contrast to ‘non-suppressive’ commensal microbes that cannot suppress MTI. Now, it has emerged that the presence of immune-suppressive microbes in the plant microbiota can suppress part of the host immune response, while favoring the colonization of opportunistic pathogens – infectious microorganisms that are normally harmless to their host but can cause disease when favorable conditions arise.
To first assess the ability of plant root commensal bacteria to interfere with defence-associated root growth inhibition (RGI), first-author Ka-Wai Ma and colleagues made use of a systematic collection of bacterial strains isolated from the roots of Arabidopsis thaliana. The authors demonstrated that after three weeks of co-culturing the bacteria with Arabidopsis thaliana in the presence of flg22 – one of the most studied microbe-derived molecules known to induce plant immune responses – 41% of the bacterial strains in the collection were able to interfere with RGI. This ability did not seem to be specific to a certain type of bacterium in the plant microbiome, as it was detected across the whole spectrum of plant microbiota associated bacteria, including Actinobacteria, Proteobacteria, Bacteroidetes and Firmicutes.
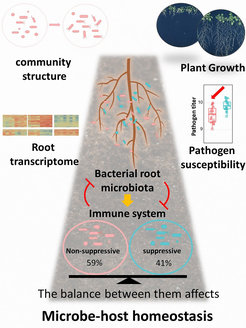
Fig. 1. Coordination of microbe-host homeostasis by crosstalk with plant innate immunity. Plants grown in soil are colonized by diverse microbes collectively known as the microbiota. Non-suppressive (red) and suppressive microbiota members (blue) exhibit contrasting capacities to modulate plant immune responses. Their combined interactions have multifaceted impacts on community structure, plant growth, expression of root genes and disease susceptibility to pathogens.
The investigators then examined the effect of synthetic communities (SynComs) made up exclusively of immune-suppressive or non-suppressive bacteria on host plants by evaluating flg22-triggered RGI and gene expression, but also the response of the SynComs themselves as a result of plant immune response stimulation by flg22. Intriguingly, they found that while SynComs made up exclusively of immune-suppressive bacteria impeded RGI, SynComs composed of non-suppressive bacteria did not. The scientists then demonstrated that activation of the host plant immune response by flg22 altered the root microbiota composition in plants colonized by non-suppressive SynComs, a finding that was not observed in root microbiota made up of immune-suppressive bacteria or a mixture of both. Furthermore, analysis of plants colonized exclusively by immune-suppressive SynComs led to altered gene expression in the host, specifically, downregulation of a subset of immune-related genes, and increased susceptibility to tested opportunistic plant pathogens. Taken together, these results indicate that suppressive and non-suppressive bacterial commensals modulate the host response in different ways primarily by promoting or impeding MTI responses, and that a balance between the two types of bacterial strains needs to be achieved in order to maintain plant homeostasis (Figure 1).
The authors’ findings strongly suggest that in order to achieve a healthy balance between plant growth and plant defence, and therefore maintain microbe-plant homeostasis, the plant root microbiota needs to contain both immune-suppressive and non-suppressive bacterial strains in balanced proportions. Having too many or not enough of either type of bacteria in the plant microbiota could be detrimental to plants, as it could result in increased disease susceptibility and poor plant growth. According to first author Ka-Wai Ma: “Although plants are known to have an intimate association with their microbiota, surprisingly, we still do not have a full understanding of their influence on the plant immune system. This study serves as a good example of how a balanced microbiota is important to modulate plant traits of interest. From a translational perspective, these findings have potential if one can manipulate the microbiota in such a way to tip the balance for the benefit of the plants.”
