Direct capture of a potent bacterial virulence activity by a plant immune receptor pair
Microbial pathogens have evolved ingenious strategies to colonize hosts, often resulting in disease. The host, in turn, can produce new proteins (receptors or antibodies) that recognize pathogen molecules to trigger defense. The debilitating effects of disease on human health and the importance of a strong and adaptable immune system to fight off infection are well known. In plants, interception of pathogens by an effective immune system is just as crucial for survival, growth and productivity. Plants rely entirely on their genetic resistance capability (innate immunity) conferred by a family of receptors expressed in individual cells. These receptors remain inactive until triggered by the actions of specific pathogen molecules which often target host basal defence pathways to promote infection. The speed with which microbial populations can produce new infection molecules (called virulence factors or effectors) places enormous pressure on plant hosts to fight back with genetically new or altered receptor recognition modes. In this article published in Cell, Le Roux et al. (2015) describe an exquisite recognition mechanism in the model plant species Arabidopsis in which a broad-sweep bacterial effector activity for promoting disease is turned into a rapid immune response.
The study focusses on a pair of Arabidopsis intracellular nucleotide-binding/leucine-rich-repeat (NLR) receptors, RRS1-R with RPS4, conferring resistance to different pathogens, including the root-infecting bacterial species, Ralstonia solanacearum, which causes an extremely costly disease in certain crops. R. solanacearum bacteria deliver a potent effector protein (PopP2) which has an acetyltransferase activity to the plant host cells. The authors show that a key target of the PopP2 acetyltransferase activity is a lysine within the conserved DNA-binding domain of a large and important family of defensive ‘WRKY’ transcription factors. In the absence of RRS1-R/RPS4 recognition, lysine-acetylation of multiple WRKY transcription factors by PopP2 dislodges them from their DNA-binding sites and disables their trans-activating functions needed for defense gene expression. This essentially dampens host basal resistance, allowing spread of the pathogen inside tissues which then leads to severe wilting of the infected plants.
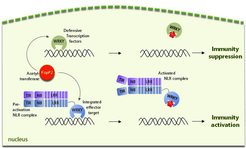
Direct capture of a pathogen defense-suppressing activity by a plant NLR pair at the DNA triggers immunity
Many genetic lines of Arabidopsis possess the PopP2-recognizing RRS1-R/RPS4 NLR pair. RRS1-R, but not RPS4, contains at its C-terminus a WRKY transcription factor DNA-binding domain. Le Roux et al (2015) show that PopP2 also directly acetylates RRS1-R, causing it to disengage from the DNA. This PopP2 action on RRS1-R likely creates a conformational change in the RRS1-R/RPS4 complex that leads to immunity activation. Thus, RRS1-R serves as both an inhibiting molecule of the NLR pair at the DNA when the pathogen effector is not present and a direct sensor of PopP2 virulence activity upon infection. The WRKY domain in RRS1-R can be viewed as an effector target ‘decoy’ which betrays the resistance-suppressing actions of PopP2 on its operational targets, the defensive WRKY transcription factors. Molecular integration of a WRKY decoy domain within the RRS1-R/RPS4 receptor complex creates an effective ‘radar’ for a powerful pathogen virulence activity which cannot be easily dispensed with by the pathogen. The observed fusion of further potential effector target decoy domains with NLR receptors in different plant species suggests a fundamental mechanism in plants for increasing receptor recognition ‘space’ as well as new avenues for engineering receptors that could effectively intercept disease-promoting activities of important crop pathogens.
