Reproductive development and the evolution of perennial life history
1. Overview
We study Arabidopsis thaliana as a model system to define the mechanisms that regulate the onset of reproduction in plants, how these are controlled by environmental cues, and how they are modified by life history. The initiation of reproduction, termed floral induction, is tightly regulated in plants to optimise their survival in fluctuating environmental conditions. Floral induction is usually triggered by seasonal cues such as daylength or winter temperatures, which cause the shoot apical meristem to produce flowers instead of leaves. This change in meristem identity and function involves complex genetic reprogramming, as well as morphological changes such as an increase in meristem size and shape. These also influence the determinacy of the inflorescence meristem, which defines the number of flowers it ultimately produces.
In addition to the mechanisms that control floral induction, which differ among species, the growth and development of floral buds can be regulated in some perennial species, so that they initiate in autumn but then arrest and do not form mature flowers until exposed to appropriate environmental cues the following spring. In this way, floral induction can be separated from the appearance of mature flowers and reproduction. Therefore, we have developed Arabis alpina, a relative of A. thaliana, as a model system to study the regulation of reproduction in the context of the perennial life-cycle.
2. Environmental and endogenous control of floral induction in Arabidopsis thaliana
A major focus of our research is to integrate the genetic network that regulates floral transition in response to environmental cues, with cellular changes that occur at the SAM. During floral transition, the SAM changes from a flat structure characteristic of the vegetative phase, to a domed shape. This phenomenon is conserved among plant species and might contribute to the indeterminacy of the inflorescence meristem by increasing SAM size and allowing more flowers to be produced. We have developed approaches to visualize morphological changes in the SAM during floral transition using confocal microscopy coupled with high-quality imaging and segmentation programs, particularly MorphoGraphX. This suite of techniques can visualize cells in different regions of the meristem during floral induction at unprecedented spatiotemporal resolution and has shown that both SAM cell size and number increase during doming, highlighting the importance of cell-cycle regulation during this process. Doming is also affected by dynamic modulation of gibberellin biosynthesis and catabolism enzymes in the SAM, which also involves transcription factors that regulate flowering time, such as SOC1 and SHORT VEGETATIVE PHASE. We are establishing a temporal and spatial framework for integrating the genes that affect flowering time with cellular changes at the SAM. One of these genes encodes the transcription factor APETALA2 (AP2), and is expressed in the SAM during vegetative growth, and downregulated at the floral transition. Loss-of-function ap2 alleles cause early flowering and reduced meristem size, whereas gain of AP2 function, due to transgenic overexpression or alleles resistant to miR172, confer delayed floral transition and a larger SAM. Therefore, AP2 function potentially links floral transition and meristem size.
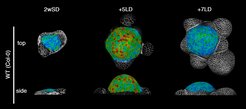

Floral transition involves dynamic developmental changes in shoot apex morphology, which can be imaged and analysed at the cellular level by segmentation programmes such as MorphoGraphX.
In collaboration with the group of Pau Formosa-Jordan, we are applying multicellular modelling to understand how the morphological changes at the shoot apex during floral transition are driven by the cell- and tissue-specific patterns of gene expression.
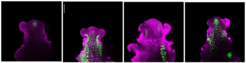
Confocal images of Arabidopsis shoot apices at floral transition in long days, showing MIRNA172A, B, C or D expression (green; left to right).
Floral induction is initiated in response to several environmental cues as well as endogenous developmental signals. Therefore, we define the gene regulatory networks that are activated by the environment and induce the morphological changes at the shoot apical meristem described above. In common with many other species, Arabidopsis initiates flowering in response to daylength via the photoperiodic pathway. This involves sensing of daylength in the leaves. Exposure of plants to long days causes accumulation of the CONSTANS protein in the leaves, which promotes transcription of the FLOWERING LOCUS T (FT) gene. The small encoded FT gene product is then translocated to the shoot apical meristem (SAM) through the phloem, where it forms a complex with the FD protein. This complex then activates a regulatory gene cascade that confers inflorescence meristem identity to the SAM and promotes the production of flowers. FD can also interact with TERMINAL FLOWER1 (TFL1), which is a protein related to FT that antagonizes the expression of downstream meristem identity genes. We are studying the mechanisms by which FT and TFL1 induce changes in shape of the shoot meristem and floral development, as well as the broader functions of FD within the SAM and stem. In collaboration with Franziska Turck, we consider how these functions are influenced by FT transcriptional regulation.
Although Arabidopsis is induced to flower by long days, it also eventually flowers under short days, when the CO–FT pathway is inactive. In short-day conditions, flowering is regulated by the endogenous, or age-related flowering pathway. As plants age, the level of microRNA156 gradually decreases, which causes an increase in the level of SQUAMOSA BINDING PROTEIN-LIKE (SPL) 15 mRNA in the SAM. We are studying how this pattern of microRNA156 expression is generated and how it contributes to SPL15 regulation. We showed that SPL15 directly activates FRUITFULL (FUL) and MIR172b, which promote flowering and inflorescence development. The transcriptional activation function of SPL15 involves interaction with other transcription factors expressed in the inflorescence meristem, such as SUPPRESSOR OF OVEREXPRESSION OF CONSTANS 1 (SOC1). We are studying how SPL15 and FUL are temporally and spatially regulated in the shoot meristem, and are identifying the functions of their target genes in conferring meristem identity. The abundance of the AP2 floral repressor mentioned above is reduced by several isoforms of miRNA172, which leads to the promotion of flowering. We are currently identifying which miRNA172 isoforms are important for the modulation of AP2 expression and how this contributes to dynamic changes in shape of the shoot meristem.
Selected recent papers
Ó’Maoiléidigh, D.S., van Driel, A., Singh, A., Sang, Q., Le Bec, N., Vincent, C., Romera-Branchat, M., Severing, E., Rafael Martinez-Gallegos R., Coupland, G. (2021). Analysis of the MIR172 family defines transcriptional and post-transcriptional mechanisms that coordinately regulate APETALA2 to control floral transition of Arabidopsis. PLoS Biology, doi:10.1371/journal.pbio.3001043.
Kinoshita, A., Vayssières, A., Richter, R., Sang, Q., Roggen, A., van Driel A.D., Smith, R.S., Coupland, G. (2020). Regulation of shoot meristem shape by photoperiodic signaling and phytohormones during floral induction of Arabidopsis. eLife 9:e60661. doi:10.7554/eLife.60661.
Madrid, E., Chandler, J.W., Coupland, G. (2020) Gene regulatory networks controlled by FLOWERING LOCUS C that confer variation in seasonal flowering and life history. Journal of Experimental Biology 72(1):4–14. doi:10.1093/jxb/eraa216.
Romera-Branchat, M., Severing, E., Pocard, C., Ohr, H., Vincent, C., Née, G., Martinez-Gallegos, R., Jang, S., Andrés, F., Madrigal, P., Coupland, G (2020) Functional Divergence of the Arabidopsis Florigen-Interacting bZIP Transcription Factors FD and FDP. Cell Reports 32(3):107966. doi:10.1016/j.celrep.2020.107966.
Sang, Q., Pajoro, A., Sun, H., Song, B., Yang, X., Stolze, S.C., Andrés, F., Schneeberger K., Nakagami, H., Coupland G. (2020) Mutagenesis of a Quintuple Mutant Impaired in Environmental Responses Reveals Roles for CHROMATIN REMODELING4 in the Arabidopsis Floral Transition. Plant Cell 32(5):1479–1500. doi:10.1105/tpc.19.00992.
Andrés, F., Kinoshita, A., Kalluri, N., Fernández, V., Falavigna, V.S., Cruz, T.M.D., Jang, S., Chiba, Y., Seo, M., Mettler-Altmann, T., Huettel, B., Coupland G. (2020) The sugar transporter SWEET10 acts downstream of FLOWERING LOCUS T during floral transition of Arabidopsis thaliana. BMC Plant Biology 20(1):53. doi:10.1186/s12870-020-2266-0.
3. Arabis as a model system to study life-history evolution and flowering patterns in perennials

The life histories of annual and perennial plants differ: annuals flower once in their life cycle and the whole plant then senesces, whereas perennials survive for many years and intersperse periods of vegetative growth and flowering, and senescence is restricted to particular plant tissues after flowering. Furthermore, perennials store nutrients at the end of the growth season in vegetative tissues that are reallocated the following year, whereas annuals transfer all nutrients to seeds.
We have developed Arabis alpina as a model system to study the control of flowering in perennials and to compare it with that in A. thaliana. We have focused on perennial-specific traits in A. alpina that limit the duration of flowering and restore vegetative growth, and define the age at which plants become sensitive to environmental cues to flower. We showed that the FLC orthologue, PERPETUAL FLOWERING 1, is differentially regulated in perennial compared with annual Brassicaceae species and terminates a reproductive episode in perennials. In addition, the age-related competence to flower is critical in perennials because it allows the plant to accumulate sufficient biomass for vegetative growth after flowering. We showed that miR156 and its target mRNAs that encode SPL transcription factors confer the time at which perennial A. alpina becomes sensitive to winter temperatures to induce flowering. We are using CRISPR-cas9 to target specific SPL genes to understand better how this process is controlled in perennials.
In collaboration with the group of Andrea Fulgione, we are studying variation in flowering responses among natural accessions of A. alpina distributed across its European range to determine how important this variation is for adaptation to different environments. In some natural populations, flowering and reproduction are mainly controlled by regulation of bud dormancy rather than floral induction. In one such Norwegian population, we are dissecting this using QTL mapping to identify the loci that distinguish early- and late-flowering individuals based on differences in the timing of bud outgrowth. To identify the genes that underlie the differences in reproductive behaviour between annuals and perennials, we have sequenced and assembled the genome of A. alpina and are sequencing the genomes of its annual relatives.
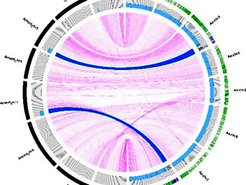
Resequencing of introgression lines carrying segments of the annual A. montbretiana genome in perennial A. alpina. Two segments (blue arcs) of the A. montbretiana genome (black outer ring) are present in the A. alpina genome (green outer ring) and were delimited by Illumina resequencing.
In another approach to identify genes that underlie annual and perennial flowering behaviour, we crossed annual A. montbretiana and perennial A. alpina to generate introgression lines carrying segments of the A. montbretiana genome in A. alpina. We identified genomic segments of A. montbretiana that conferred extreme late flowering or early flowering to A. alpina. These segments contain genes encoding MADS-box transcriptions that are either duplicated in the annual accession or are differentially transcribed in the annual compared with the perennial. We are studying the functions of these genes in both species, and characterizing other introgressions that alter reproductive behavior of A. alpina.
Selected recent papers
Madrid, E., Chandler, J.W., Coupland, G. (2020) Gene regulatory networks controlled by FLOWERING LOCUS C that confer variation in seasonal flowering and life history. Journal of Experimental Biology 72(1):4–14. doi:10.1093/jxb/eraa216.
Hyun, Y., Vincent, C., Tilmes, V., Bergonzi, S., Kiefer, C., Richter, R., Martinez-Gallegos, R., Severing, E., Coupland, G. (2019) A regulatory circuit conferring varied flowering response to cold in annual and perennial plants. Science 363(6425):409–412. doi:10.1126/science.aau8197.
Laenen, B., Tedder, A., Nowak, M.D., Toräng, P., Wunder, J., Wötzel, S., Steige, K.A., Kourmpetis, Y., Odong, T., Drouzas, A.D., Bink, M.C.A.M., Ågren, J., Coupland, G., Slotte, T. (2018) Demography and mating system shape the genome-wide impact of purifying selection in Arabis alpina. Proceedings of the National Academy of Science of the United States of America. 115(4):816–821. doi.org/10.1073/pnas.1707492115.
Kiefer, C., Severing, E., Karl, R., Bergonzi, S., Koch, M., Tresch, A., Coupland, G. (2017). Divergence of annual and perennial species in the Brassicaceae and the contribution of cis-acting variation at FLC orthologues. Molecular Ecology 26:3437–3457. doi:10.1111/mec.14084.
Mateos, J. L., Tilmes, V., Madrigal, P., Severing, E., Richter, R., Rijkenberg, C.W.M., Krajewski, P., Coupland, G. (2017). Divergence of regulatory networks governed by the orthologous transcription factors FLC and PEP1 in Brassicaceae species. Proceedings of the National Academy of Sciences of the United States of America 114(51):E11037–E11046. doi:10.1073/pnas.1618075114.
4. More on our work
The above is a summary of some aspects of our most recent work. A full list of publications is available on Google Scholar
Most of our publications are available as PDFs for download on ResearchGate at the link https://www.researchgate.net/profile/George_Coupland/contributions
If you would like more information on our work or are interested in joining us as an Intern, Masters student, PhD student or Post doc, please contact George Coupland by Email.



