Colletotrichum higginsianum Genome Project Description
Data Release Statement
The Colletotrichum higginsianum genome has been sequenced at the Max Planck Institute for Plant Breeding Research, Cologne with funds from the Max Planck Society. We intend to publish the complete annotated genome in a peer-reviewed journal as soon as possible. The permission of the principal investigator Dr Richard O’Connell must be obtained before publishing any genome-scale analyses based on unpublished sequences, genes or other features presented on this web site. In any publications, users of this resource are requested to acknowledge the Max Planck Institute for Plant Breeding Research and to cite the database as follows:
Max Planck Institute for Plant Breeding Research Colletotrichum higginsianum Genome Project, http://www.mpipz.mpg.de/colletotrichum_higginsianum_genome_project_description
Questions concerning this project or use of the data should be sent to Emiel Ver Loren van Themaat or Dr Richard O'Connell
The Colletotrichum higginsianum-Arabidopsis thaliana pathosystem
Colletotrichum is a large genus of haploid Ascomycete fungi, comprising approximately 20 species, which cause destructive anthracnose diseases on many agricultural and horticultural crops throughout the world. The sexual (teleomorphic) stage is Glomerella (family Glomerellaceae, class Sordariomycetes), but a sexual stage has not been reported for some species, including Colletotrichum higginsianum.
Colletotrichum higginsianum causes anthracnose leaf spot disease on many cultivated forms of Brassica and Raphanus, but can also infect Arabidopsis thaliana. This provides an attractive model pathosystem for dissecting fungal pathogenicity and plant resistance, in which both partners can be genetically manipulated. Thus, C. higginsianum can be cultured axenically and stably transformed, allowing random mutagenesis and targeted gene disruption, over-expression, gene silencing and protein-tagging.
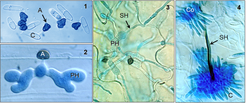
The pathogen employs a hemibiotrophic infection strategy to invade host plants, involving differentiation of a series of specialised cell types (infection structures). After initial penetration of host epidermal cells by appressoria, the fungus grows biotrophically inside living epidermal cells, producing bulbous primary hyphae that invaginate the host plasma membrane, before later switching to a destructive necrotrophic phase associated with filamentous secondary hyphae. The fungus completes its asexual cycle by producing sporulating structures called acervuli on the surface of the dead tissue.
Phylogenetic analysis based on sequencing the ITS regions of rDNA indicates that C. higginsianum forms part of a group of closely-related taxa that also includes C. destructivum (tobacco and legume pathogen) and C. linicola (flax pathogen). A characteristic feature of all three species is that the initial biotrophic phase of infection is restricted to a single host epidermal cell, in contrast to other hemibiotrophic Colletotrichum species which establish biotrophy in many host cells.
The genome sequencing project
The aim of this project is to produce a high-quality reference genome assembly for C. higginsianum, which will provide a valuable resource for:
- studying mechanisms of fungal pathogenicity
- identification of secreted effector proteins required for host manipulation
- comparative genomic analysis of the evolutionary and functional relationships between hemibiotrophy and other pathogenic lifestyl
The genome of the maize pathogen, C. graminicola, has already been sequenced at the Broad Institute and a high quality draft assembly has been released:
Colletotrichum graminicola database
This provides the opportunity to compare the genomes of two closely-related species which have contrasting hemibiotrophic lifestyles and also differ in their host specificity. It will also enable the identification of genes undergoing rapid evolution (diversifying selection), which are likely to be involved in interactions with the host plant, e.g. those encoding effector proteins. Overall, we envisage that C. graminicola will provide a model for anthracnose diseases on monocot hosts, while C. higginsianum will become the model of choice for studying Colletotrichum infection of dicot plants.
Strategy used for whole-genome sequencing
For sequencing the estimated 50 Mbp genome of C. higginsianum, we have used a combination of next-generation sequencing technologies. Sequencing and assembly were conducted by GATC Biotech AG (Konstanz, Germany). The following raw sequence data have been generated:
| Roche 454 Titanium shot-gun reads | 1.22 Gb | 24X |
| Sanger fosmid end sequences (900 clones) | 870 kb | 0.2X |
| Illumina paired-end (200bp inserts, 2 x 76 bp reads) | 1.22 Gb | 24X |
| Illumina paired-end (3 kb inserts, 2 x 36 bp reads) | 0.93 Gb | 19 |
| Illumina paired-end (10 kb inserts, 2 x 76 bp reads) | 1.5 Gb | 30X |
The raw sequence reads will be made publically available through the NCBI trace file database.
Assembly statistics
The first draft genome assembly is based on 1.22 Gbp of 454-Titanium shot-gun sequencing data assembled using the Roche Newbler programme.
| Total number of contigs: | 8,303 |
| Total length of contigs: | 47.15 Mb |
| Contigs >1kb: | 5,420 |
| N50 length of contigs >1kb: | 14.78 kb |
| Raw data in contigs >1kb: | 98% |
Out of 248 core genes expected to be found in all eukaryotic genomes [Parra et al. (2009) Nucl. Acids Res. 37, 289–297], 236 are present in this assembly, suggesting that 95% of the total gene space is already covered. Further assembly is ongoing and future updates of the assembly and annotation will be made publically available through this website and the Broad Institute’s Fungal Genomics website.
Annotation of the genome is currently performed in collaboration with Dr Li-Jun Ma at the Broad Institute and the annotated genome is planned for release in summer 2010.
Searching the sequence data
The genome browser presents all 8,303 genomic contigs from the current assembly. Gene models predicted using three different programmes are displayed:
- Conrad (trained with Fusarium ESTs): 19,400 predicted ORFs
- FGenesH (Magnaporthe matrix): 15,900 ORFs
- FGenesH (Fusarium matrix): 12,700 ORFs
- Genemark (ab initio gene calling): 15,734 ORFs
The Gbrowse database also displays:
- results of BLASTX homology searches against the NCBI non-redundant protein database
- sequences homologous to the CEGMA set of core eukaryotic genes [Parra et al. (2009) Nucl. Acids Res. 37, 289–297]
- regions of repetitive DNA
- regions showing evidence of repeat-induced point mutation (RIP)
The sequence data can be searched using a local BLAST server, which links the results of homology searches to the corresponding genomic contigs.
Project team at the Max Planck Institute for Plant Breeding Research, Köln
Principal Investigators: Dr Richard O’Connell and Emiel Ver Loren van Themaat, Plant-Microbe Interactions Department
Dr Kurt Stüber, Bioinformatics Services Group
Dr Richard Reinhardt, MPIZ Genome Centre
Dr Heiko Schoof, Plant Computational Biology Group
Collaborators
Dr Lisa Vaillancourt, University of Kentucky, Lexington, KY, USA
Dr Mike Thon, CIALE, University of Salamanca, Salamanca, Spain
Dr Li-Jun Ma, Broad Institute of MIT, Cambridge, MA, USA
Prof Ken Shirasu, RIKEN Plant Science Center, Yokohama, Japan
Prof. Yasuyuki Kubo, Kyoto Prefectural University, Kyoto, Japan
Dr Yoshitaka Takano, Kyoto University, Kyoto, Japan
Service Providers
GATC Biotech AG, Konstanz, Germany (genome sequencing and assembly)
DNAStar Lasergene (genome assembly and scaffolding)
