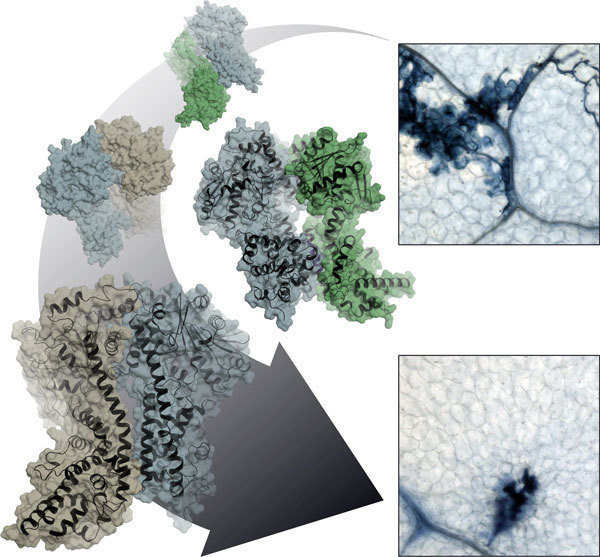
Molecular snapshot of the plant immune system’s signal box
The molecular architecture of three key proteins and their complexes reveals how plants fine-tune their immune response to pathogens
Plants rarely get sick in their natural environment. When the threat of infection arises, a quick decision is made about the necessary countermeasures. The course is set by a protein which forms complexes with its partner proteins for this purpose. Jane Parker from the Max Planck Institute for Plant Breeding Research in Cologne and Karsten Niefind from the Institute of Biochemistry at the University of Cologne have, together with colleagues, determined the three-dimensional structure of such a key complex. These molecular close-ups will enable scientists to reach a better understanding of plant immunity in the near future.
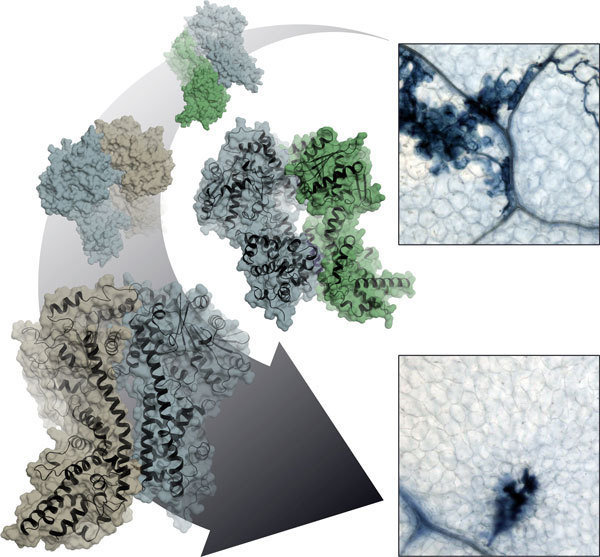
Like other organisms, plants defend themselves against attack from disease-causing microbes. Because these immune responses are associated with considerable stress and energy expenditure, the infected cells make very specific decisions about the actions actually required. To do this, they employ several layers of defence which come into force at different stages of pathogen infection, gradually ramping up the host immune response. Sometimes, this relies on a radical solution at the site of the infection and the plant sacrifices the infected cell by programmed cell death. Death, however, must be meticulously controlled. A rapidly administered ‘acute’ local defence response also usefully primes the remainder of the plant against subsequent attack.
“Effector-triggered immunity” is one such line of defence. Here, plant immune receptors inside the cell recognise specific disease factors produced by a highly adapted pathogen. What happens after that depends on which complexes the mobile protein EDS1, moving between the cell nucleus and cytoplasm, forms with its partner proteins, PAD4 and SAG101. EDS1 sets the course of the immune response by forming distinct complexes – known as heterodimeric complexes – with its partners. The immune response varies according to the nature of the complex formed. The three proteins are structurally similar and the nature of each heterodimeric complex likely dictates the resistance output. “We would like to know how the plant resistance signalling system works and makes decisions in a dynamic way to confer resistance. Handling pathogen stress likely involves exquisite communication between different pathways ’” says Jane Parker, explaining her interest in the three proteins.
Together with Jane Parker and Johannes Stuttmann from the Max Planck Institute for Plant Breeding Research and other colleagues, Karsten Niefind and Stephan Wagner from the University of Cologne determined the atomic structure of the EDS1-SAG101 complex by means of protein x-ray crystallography, and extrapolated a model for the EDS1-PAD4 complex from this three-dimensional structure. All three proteins have a domain at their N terminals with a α/β hydrolase fold. This fold is one of the most successful architectures in protein evolution. It also arises in a similar form in fat-splitting lipases and various other enzymes. “Naturally, we wanted to know whether EDS1 has a lipase activity on its own or in the heterodimeric complex, and whether EDS1 needs this lipase activity for its functions,” says Parker. “Surprisingly, the structure revealed to us that this is not the case, as the potentially active centre is completely shielded by a kind of cover. We also could not detect any lipase activity in the test tube.” What’s more: Parker and her colleagues showed that Arabidopsis plants, in which the putative catalytic centre of both EDS1 and PAD4 has been destroyed through mutations, are nonetheless fully competent in resistance to certain pathogens as the wild type proteins.
So how does EDS1 accomplish its resistance signalling task, if not through catalytic activity? The x-ray structure of the EDS1/SAG101 heterodimer complex provides a valuable pointer here. EDS1 has a conspicuous protuberance or loop in its N terminal domain which slips into a matching cavity in SAG101 like a key in a lock. PAD4 has a similar cavity and can accommodate the same EDS1 loop. The Cologne-based scientists engineered mutations into these regions and in this way showed that the central task of the lipase-like domains is likely in the provision of these contact sites. Only when the cavity and protuberance are matched can the important C-terminal domains of both complex partners be brought into spatial proximity to create a new functional module. Without the lipase-like domains, there are no stable complexes, and without heterodimerised C-terminals there is no immune response!
Intensive investigations are underway to find out how formation of the different EDS1 heterodimers in the cell proceeds and the precise role of each exclusive heterodimer. “The x-ray structure not only revealed to us the mechanism of heterodimerisation in the EDS1 protein family, it also shows us prominent surfaces and binding pockets which only exist in the dimers and will likely provide the key to understanding this important plant immunity mechanism,” explains Niefind.